“Visual mining of neuro-metaspaces” by Joshi, Bowman, Hasson, Liu, Toga, et al. …
Conference:
Type(s):
Title:
- Visual mining of neuro-metaspaces
Presenter(s)/Author(s):
Abstract:
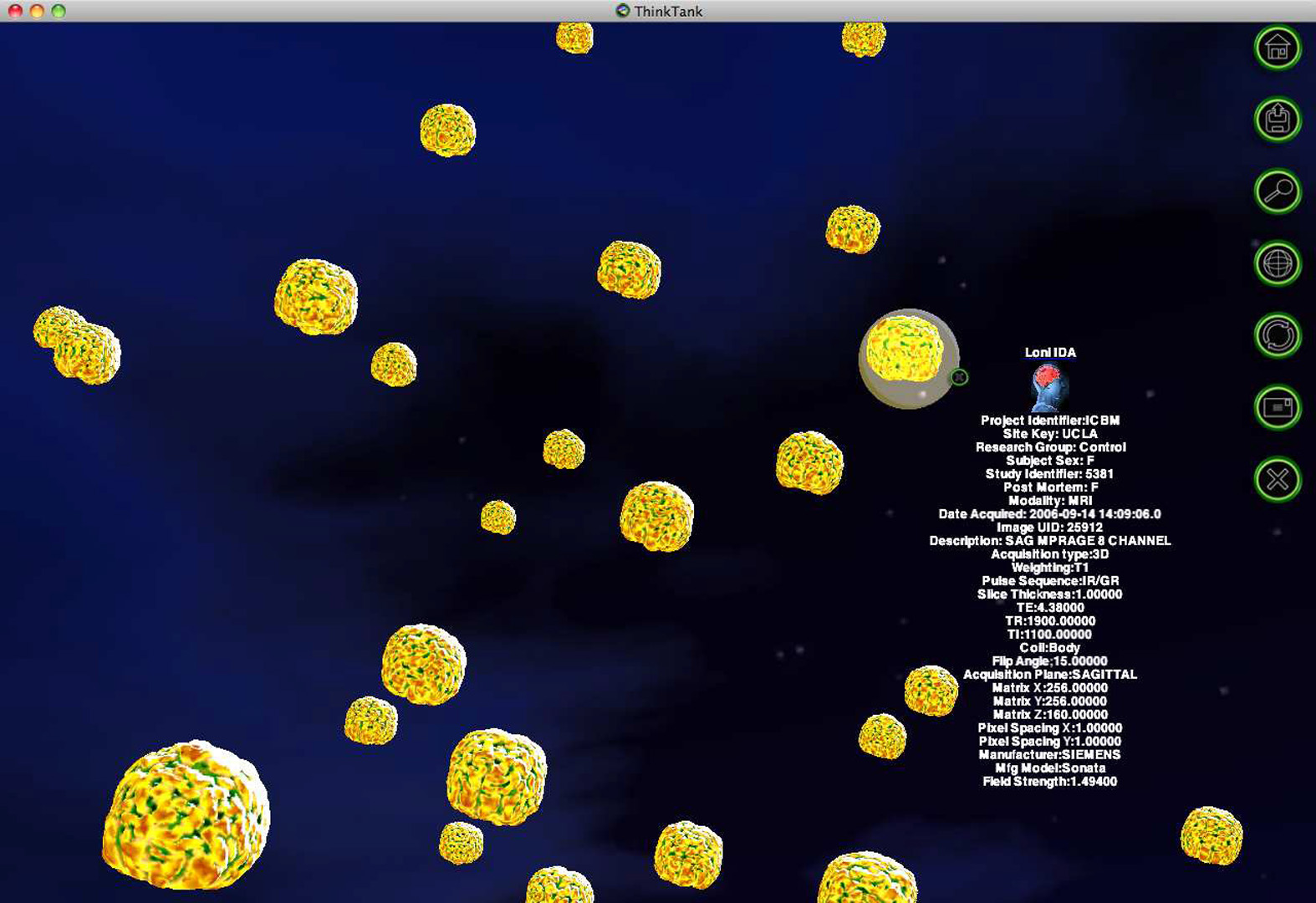
Large scale neuroimaging data archival protocols are gradually becoming ubiquitous in both research as well as clinical settings. Current user-database interfaces are limited to textual searches and often require data-specific knowledge for performing queries. This is proving to be an obstacle for researchers who wish to obtain a holistic view of the data before designing pilot neuroscientific studies or even formulating statistical hypotheses. Instead of providing a restricted, unidimensional view of the data, we seek to place a multi-dimensional view of the entire neurodatabase at the user’s disposal. With the aim of visual navigation of complete neuro-repositories, we introduce the concept of brain meta-spaces. The meta-space models the implicit nonlinear manifold where the neurological data resides, and encodes pair-wise dissimilarities between all individuals in a population. Additionally, the novelty in our approach lies in the user ability to simultaneously view and interact with many brains at once but doing so in a vast meta-space that encodes (dis)similarity in morphometry.
References:
1. Dinov, I., Van Horn, J. D., and et al. 2009. Efficient, distributed and interactive neuroimaging data analysis using the loni pipeline. Frontiers in Neuroinformatics, 38.
2. Joshi, S. H., Van Horn, J., and Toga, A. W. 2009. Interactive exploration of neuroanatomical meta-spaces. Frontiers in Neuroinformatics 3, 38.